Haowen Zhou wasn’t looking for a breakthrough. He was debugging. Somewhere in the code for an unrelated project, something wasn’t adding up, and he and his colleague Shi “Josh” Zhao were methodically working through the numbers when they noticed something odd in the images. When they combined photographs taken from two slightly different illumination angles, a pattern of stripes appeared in the data. Faint, but unmistakeable. And stranger still, the stripes changed depending on how blurry the image was. “If we defocused more, the fringe would be much denser,” Zhou says. “If we defocused less, it would be more spread out. So, it seemed there was a strong correlation between the defocus value and the fringe density.”
What the two Caltech graduate students had stumbled onto was a new way of solving a problem that has irritated microscopists for as long as there have been microscopes: keeping the damn thing in focus.
It sounds trivial. It isn’t. Every time you move a slide under a microscope, the sample drifts out of the focal plane and the image blurs. For a scientist or clinician working through dozens or hundreds of samples in an automated imaging pipeline, that constant refocusing is a serious drag. Existing autofocus systems exist, but most are slow, fiddly, or break down when faced with anything more complicated than a thin flat specimen. Three-dimensional tissue samples, organoids, thick brain slices, whole embryos. those are where existing methods tend to give up. The focus drifts, the images degrade, and someone has to intervene.
Stripes as a Ruler
The technique Zhou and Zhao developed, working in the lab of Changhuei Yang at Caltech, is called Digital Defocus Aberration Interference, or DAbI. The underlying hardware requirement is almost aggressively modest: two LEDs. The LEDs illuminate a sample from slightly different angles, and a computer captures one photograph under each source of illumination. Add those two images together in software and the interference fringe pattern appears. The denser the stripes, the further the sample is from the focal point. It’s basically using light physics as a ruler, a way of measuring exactly how far off-focus you are without ever having to squint at an image and guess.
The performance numbers are, frankly, difficult to square with the simplicity of the setup. For thin flat samples, DAbI extends the effective autofocus range to more than 400 times what the natural depth-of-field of a standard microscope lens allows. For 3D specimens up to 150 micrometers deep, it still achieves a range roughly 300 times the natural limit. No comparable technique has managed the 3D case reliably before. “This offers truly robust autofocusing of 3D samples, which has never been possible with other techniques,” Zhao says.
The system has been tested across six different microscope types, including brightfield, confocal, and widefield fluorescence setups. It worked consistently across all of them, which matters because most autofocus approaches are tuned to a specific configuration and struggle to generalize.
Brain Slices and Organoids
To push DAbI on the most challenging biological material they could find, Zhou and Zhao reached out to Viviana Gradinaru’s neuroscience lab, also at Caltech. Gradinaru’s postdoctoral researcher Yujie Fan helped prepare samples: brain slices between 50 and 100 micrometers thick, and organoids, lab-grown mini-organs with the kind of lumpy, irregular geometry that sends conventional autofocus systems into confusion. “Because of the complex 3D structures and irregular shapes of these thick samples, it’s really hard for the current microscope system to find the perfect focus automatically,” Fan explains. DAbI, she says, “performed exceptionally well even with thick biological samples.”
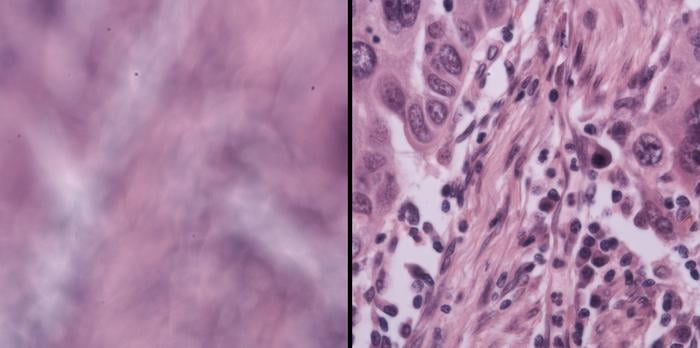
The team also tested it on live mouse embryos, algae, bacteria, and human lung cancer tissue provided by collaborators at Washington University in St. Louis. The ecological and oncological samples brought their own challenges, each a different kind of optical headache, and DAbI handled all of them. “Our technique, which is enabled by a physics-based observation, is reliable, high performance, and also very simple,” Zhou says. “This makes it useful and powerful for automated, high-throughput microscopy.”
The significance for labs doing large-scale tissue screening is hard to overstate. High-throughput imaging, the kind used in drug discovery or cancer diagnostics, depends on processing enormous numbers of samples automatically. If the autofocus fails even occasionally, data quality degrades across the entire run, sometimes in ways that aren’t obvious until analysis. A system that can reliably maintain focus through thick 3D material, without manual intervention, changes what’s actually possible in that pipeline. Fan puts it fairly directly: integrating DAbI into standard microscopes would mean “not only saving researchers like me a massive amount of time, but also making imaging-based high-throughput screening of 3D tissue possible.”
What Comes Next
Caltech filed a provisional patent on DAbI in July 2025, which suggests the inventors have one eye on commercialisation. The practical pathway isn’t complicated: the hardware addition is two LEDs and a software layer, nothing that would require retooling a microscope from scratch. Whether that simplicity is enough to get it adopted into commercial platforms remains to be seen; instrumentation companies have their own development cycles and their own ideas about what to upgrade.
Still, the serendipity of the discovery has a pleasingly old-fashioned quality. Not a product of deliberate engineering toward a stated goal, but a pattern noticed in the noise of something else entirely, then followed wherever it led. Microscopy has been bottlenecked at the focusing step for a long time. The solution, it turns out, was hiding in a debugging session.
Source: Zhou et al., Nature Communications (2026)
Frequently Asked Questions
How does DAbI actually keep a microscope in focus?
DAbI uses two LEDs to illuminate a sample from slightly different angles, then combines the resulting images in software. The combination produces a striped interference pattern whose density changes predictably depending on how far the sample is from the focal plane. A computer reads the stripe density and calculates exactly how much to adjust the focus, all without human input. The physics-based nature of the relationship is what makes it reliable across very different sample types and microscope configurations.
Why is autofocusing thick 3D samples so hard normally?
Conventional autofocus systems work well with thin, flat specimens because the focal plane is easy to locate. Thick 3D samples like brain slices or organoids have complex, irregular internal structures that scatter light unpredictably, making it genuinely difficult for standard algorithms to find where focus is sharpest. Most existing techniques either give up or require time-consuming manual adjustments at regular intervals. DAbI sidesteps this by measuring defocus directly from the interference fringe pattern rather than trying to assess image sharpness.
Could this make drug discovery faster?
Possibly, yes. High-throughput screening in drug discovery relies on imaging enormous numbers of samples automatically, and focus failures anywhere in the pipeline degrade data quality across the whole run. A system that maintains reliable focus through 3D tissue without intervention could allow labs to image samples they currently have to handle manually, or to run larger automated screens with greater confidence in the results. Whether that advantage translates into faster discovery depends on where the autofocus step is the actual bottleneck.
Is this expensive to add to an existing microscope?
The hardware requirement is minimal: two LEDs and the associated software. There’s no need to rebuild the optical path or replace the objective lens. In principle, DAbI could be retrofitted to existing instruments without significant cost, though commercial adoption depends on whether microscope manufacturers integrate it into their platforms. A provisional patent was filed in 2025, so that process is at least underway.
ScienceBlog.com has no paywalls, no sponsored content, and no agenda beyond getting the science right. Every story here is written to inform, not to impress an advertiser or push a point of view.
Good science journalism takes time — reading the papers, checking the claims, finding researchers who can put findings in context. We do that work because we think it matters.
If you find this site useful, consider supporting it with a donation. Even a few dollars a month helps keep the coverage independent and free for everyone.