Sometimes the most powerful discoveries are the ones scientists walk past for decades. Researchers studying a bacterium that has lived in lab freezers since the 1950s just found an antibiotic 100 times more potent than anything they knew the organism could make. The catch? It was hiding as a throwaway ingredient in a chemical recipe the bug had been following all along.
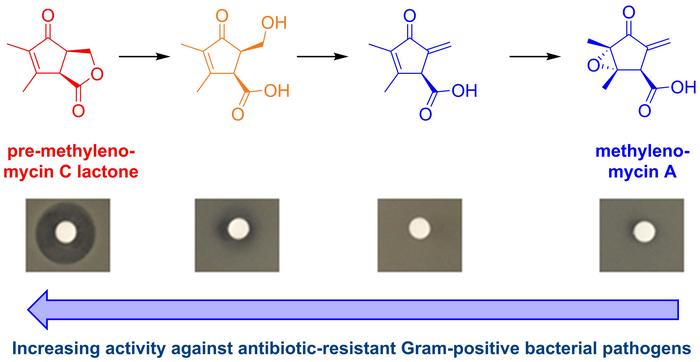
The compound, called pre-methylenomycin C lactone, is an intermediate step in the production of methylenomycin A, a known but relatively weak antibiotic first isolated 50 years ago. Chemists from the University of Warwick and Monash University discovered it by essentially breaking the bacterium’s assembly line: deleting genes one by one to see what half-finished molecules would pile up.
“Methylenomycin A was originally discovered 50 years ago and while it has been synthesized several times, no one appears to have tested the synthetic intermediates for antimicrobial activity,” says Professor Greg Challis, who co-led the study published in the Journal of the American Chemical Society. “By deleting biosynthetic genes, we discovered two previously unknown biosynthetic intermediates, both of which are much more potent antibiotics than methylenomycin A itself.”
A Familiar Bug with Unfamiliar Tricks
The bacterium in question, Streptomyces coelicolor, is something of a lab mascot. It is the kind of organism graduate students cut their teeth on, a model species for understanding how soil microbes churn out medically useful chemicals. That it still had secrets to reveal is, to put it mildly, unexpected.
Dr. Lona Alkhalaf, Assistant Professor at Warwick and co-lead author, captures the surprise: the bacterium “is a model antibiotic-producing species that’s been studied extensively since the 1950s. Finding a new antibiotic in such a familiar organism was a real surprise.” It is a bit like discovering your neighbor has been sitting on a winning lottery ticket for half a century.
The researchers suspect that S. coelicolor once made pre-methylenomycin C lactone as its primary defense compound but evolved to modify it into the weaker methylenomycin A, possibly for a different biological role. Evolution does not always optimize for maximum punch.
Potency Against the Worst Offenders
When the team tested pre-methylenomycin C lactone against a panel of bacteria, it demolished Gram-positive pathogens, including the strains behind MRSA (methicillin-resistant Staphylococcus aureus) and VRE (vancomycin-resistant Enterococcus). These are not minor players: VRE is on the WHO’s list of high-priority pathogens, and vancomycin is often a last-resort drug.
Even more striking, the researchers could not coax Enterococcus bacteria to develop resistance to the compound under conditions that routinely produce vancomycin resistance. That durability is rare and precious in the antibiotic world, where resistance can emerge faster than new drugs reach the market.
The compound’s structure is mercifully simple, which bodes well for synthesis. A Monash team led by Professor David Lupton published a scalable synthesis route earlier this year, meaning the molecule can be produced in quantities sufficient for further testing. Lupton notes that the synthesis “should enable the creation of diverse analogues that can be used to probe the structure-activity relationship and mechanism of action.”
The next phase is pre-clinical testing, the gauntlet that separates laboratory curiosities from clinical candidates. With antimicrobial resistance claiming roughly 1.1 million lives annually, the stakes are clear. Challis frames the discovery as a potential paradigm shift: “By identifying and testing intermediates in the pathways to diverse natural compounds, we may find potent new antibiotics with more resilience to resistance.”
It is a reminder that nature’s drafts are sometimes better than the final copy. The question now is whether this particular rough draft can make it all the way to the pharmacy.
Journal of the American Chemical Society: 10.1021/jacs.5c12501
ScienceBlog.com has no paywalls, no sponsored content, and no agenda beyond getting the science right. Every story here is written to inform, not to impress an advertiser or push a point of view.
Good science journalism takes time — reading the papers, checking the claims, finding researchers who can put findings in context. We do that work because we think it matters.
If you find this site useful, consider supporting it with a donation. Even a few dollars a month helps keep the coverage independent and free for everyone.